Documentation | Installation | Tutorial | API reference | GitHub | Qiita (Japanese)
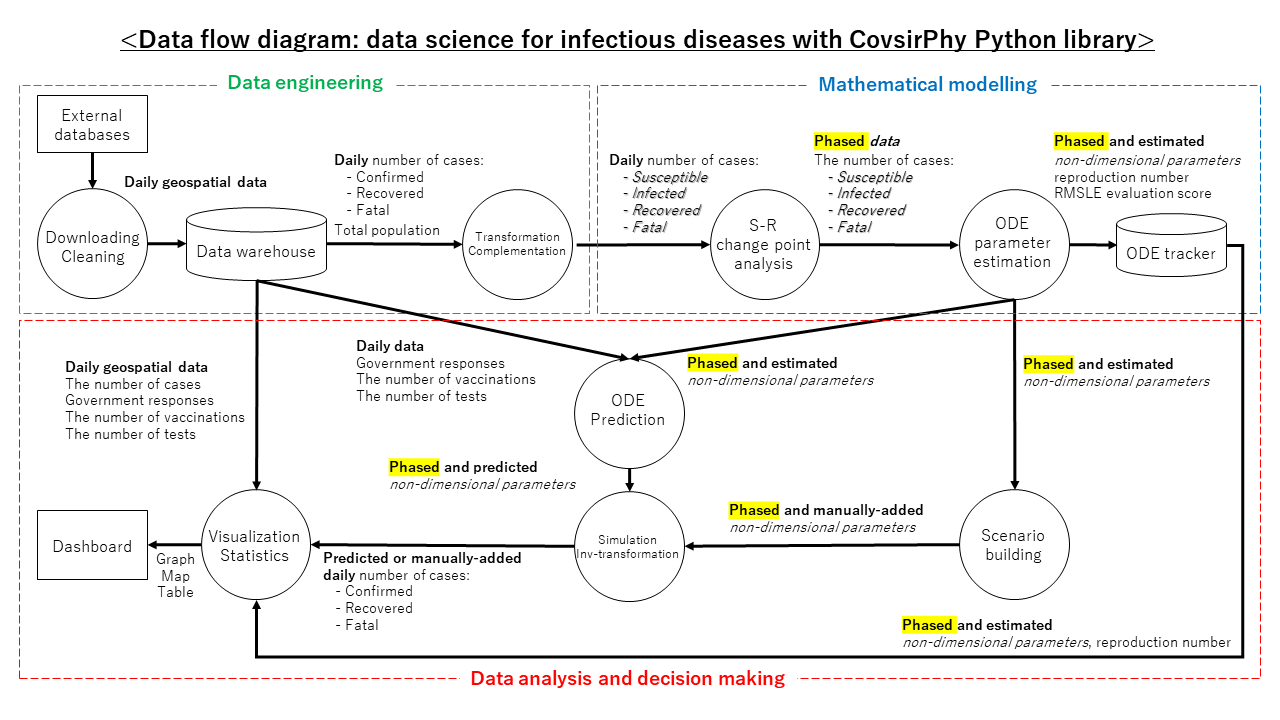
CovsirPhy is a Python library for infectious disease (COVID-19: Coronavirus disease 2019, Monkeypox 2022) data analysis with phase-dependent SIR-derived ODE models. We can download datasets and analyze them easily. Scenario analysis with CovsirPhy enables us to make data-informed decisions.
- Monitor the spread of COVID-19/Monkeypox with SIR-derived ODE models
- Predict the number of cases in each country/province
- Find the relationship of reproductive number and measures taken by each country
If you have ideas or need new functionalities, please join this project. Any suggestions with Github Issues and Twitter: @lisphilar are always welcomed. Questions are also great. Please refer to Guideline of contribution.
The latest stable version of CovsirPhy is available at PyPI (The Python Package Index): covsirphy and supports Python 3.8 or newer versions. Details are explained in Documentation: Installation.
pip install --upgrade covsirphyQuickest tour of CovsirPhy is here. The following codes analyze the records in Japan.
import covsirphy as cs
# Data preparation,time-series segmentation, parameter estimation with SIR-F model
snr = cs.ODEScenario.auto_build(geo="Japan", model=cs.SIRFModel)
# Check actual records
snr.simulate(name=None);
# Show the result of time-series segmentation
snr.to_dynamics(name="Baseline").detect();
# Perform simulation with estimated ODE parameter values
snr.simulate(name="Baseline");
# Predict ODE parameter values (30 days from the last date of actual records)
snr.build_with_template(name="Predicted", template="Baseline");
snr.predict(days=30, name="Predicted");
# Perform simulation with estimated and predicted ODE parameter values
snr.simulate(name="Predicted");
# Add a future phase to the baseline (ODE parameters will not be changed)
snr.append();
# Show created phases and ODE parameter values
snr.summary()
# Compare reproduction number of scenarios (predicted/baseline)
snr.compare_param("Rt");
# Compare simulated number of cases
snr.compare_cases("Confirmed");
# Describe representative values
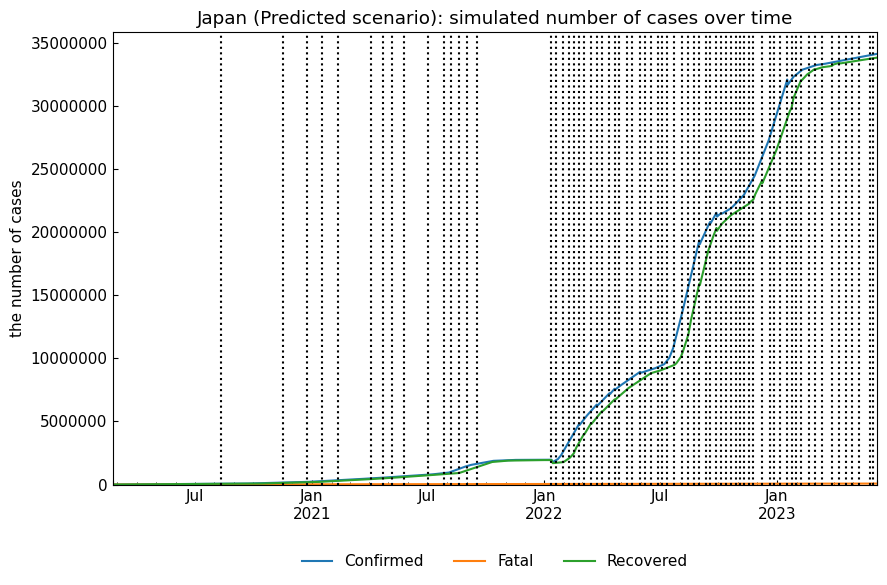
snr.describe()Output of snr.simulate(name="Predicted");
Tutorials of functionalities are included in the CovsirPhy documentation.
- Data preparation
- Data Engineering
- SIR-derived ODE models
- Phase-dependent SIR models
- Scenario analysis
- ODE parameter prediction
Release notes are here. Titles & links of issues are listed with acknowledgement.
We can see the release plan for the next stable version in milestone page of the GitHub repository. If you find a highly urgent matter, please let us know via issue page.
CovsirPhy library is developed by a community of volunteers. Please see the full list here.
This project started in Kaggle platform. Hirokazu Takaya (@lisphilar) published Kaggle Notebook: COVID-19 data with SIR model on 12Feb2020 and developed it, discussing with Kaggle community. On 07May2020, "covid19-sir" repository was created. On 10May2020, covsirphy version 1.0.0 was published in GitHub. The first release in PyPI (version 2.3.0) was on 28Jun2020. Many APIs were reviewed via 2.x series and version 3.0.0 was released on 12May2023.
Please support this project as a developer (or a backer).
Please refer to LICENSE file.
Please cite this library as follows with version number (import covsirphy as cs; cs.__version__).
Hirokazu Takaya and CovsirPhy Development Team (2020-2024), CovsirPhy version [version number]: Python library for COVID-19 analysis with phase-dependent SIR-derived ODE models, https://github.com/lisphilar/covid19-sir
This is the output of covsirphy.__citation__.
import covsirphy as cs
cs.__citation__We have no original papers the author and contributors wrote, but note that some scientific approaches, including SIR-F model, S-R change point analysis, phase-dependent approach to SIR-derived models, were developed in this project.